library(readr) # tidyverse's
library(data.table)
library(microbenchmark)
# Generating a large dataset of random numbers
set.seed(1231)
x <- runif(5e4 * 10) |> matrix(ncol = 10) |> data.frame()
# Creating tempfiles
temp_dt <- tempfile(fileext = ".csv")
temp_tv <- tempfile(fileext = ".csv")
temp_r <- tempfile(fileext = ".csv")The data.table R package
For most cases, the most proper data wrangling tool in R is the dplyr package. Nonetheless, when dealing with large amounts of data, the data.table package is the fastest alternative available for doing data processing within R (see the benchmarks).
Reading and Writing Data
Reading and writing operations with data.table’s fread and fwrite are highly optimized. Here is a benchmark we can do on our own:
bm <- microbenchmark(
readr = write_csv(x, temp_tv, num_threads = 1L, progress = FALSE),
data.table = fwrite(
x, temp_dt, verbose = FALSE, nThread = 1L,
showProgress = FALSE),
base = write.csv(x, temp_r),
times = 20
)Warning in microbenchmark(readr = write_csv(x, temp_tv, num_threads = 1L, :
less accurate nanosecond times to avoid potential integer overflowsbmUnit: milliseconds
expr min lq mean median uq max neval
readr 19.83141 24.03928 36.83226 26.99321 28.25456 240.43183 20
data.table 22.78067 25.24101 27.63036 27.15459 30.10759 32.39791 20
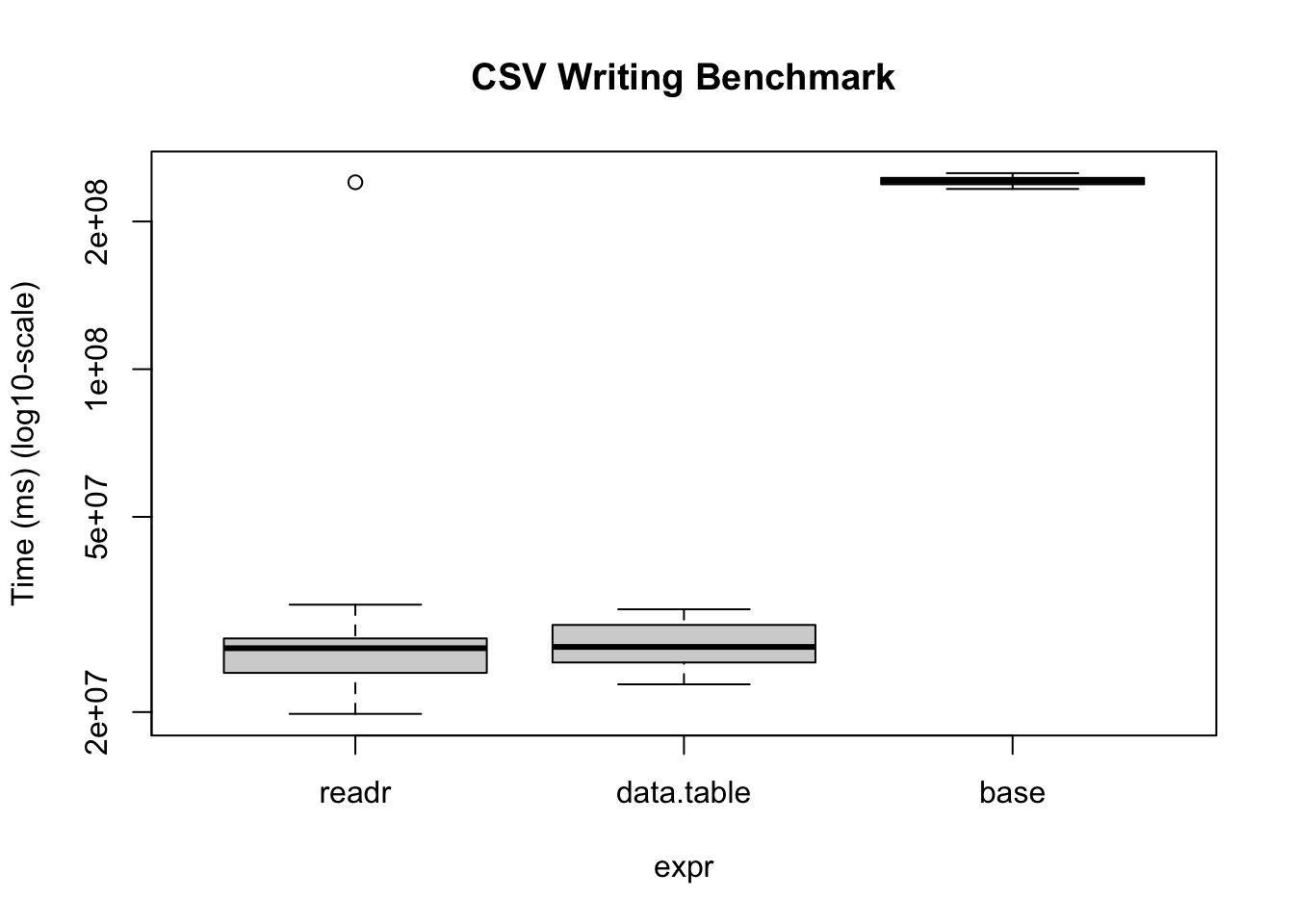
base 233.02502 238.19760 241.98725 242.07634 245.42001 250.93275 20# We can also visualize it
bm |>
plot(
log = "y",
ylab = "Time (ms) (log10-scale)",
main = "CSV Writing Benchmark"
)
The same thing applies when reading data
# Writing the data
fwrite(x, temp_r, verbose = FALSE, nThread = 1L, showProgress = FALSE)
# Benchmarking
bm <- microbenchmark(
readr = read_csv(
temp_r, progress = FALSE, num_threads = 1L,
show_col_types = FALSE
),
data.table = fread(
temp_r, verbose = FALSE, nThread = 1L,
showProgress = FALSE
),
base = read.csv(temp_r),
times = 10
)
bmUnit: milliseconds
expr min lq mean median uq max neval
readr 39.94503 41.21443 48.01355 43.21878 44.05876 97.11883 10
data.table 9.46854 9.85722 12.65471 10.50902 11.52186 22.31364 10
base 167.90447 199.18850 239.84786 217.32075 250.02190 458.34343 10bm |>
plot(
log = "y",
ylab = "Time (ms) (log10-scale)",
main = "CSV Reading Benchmark"
)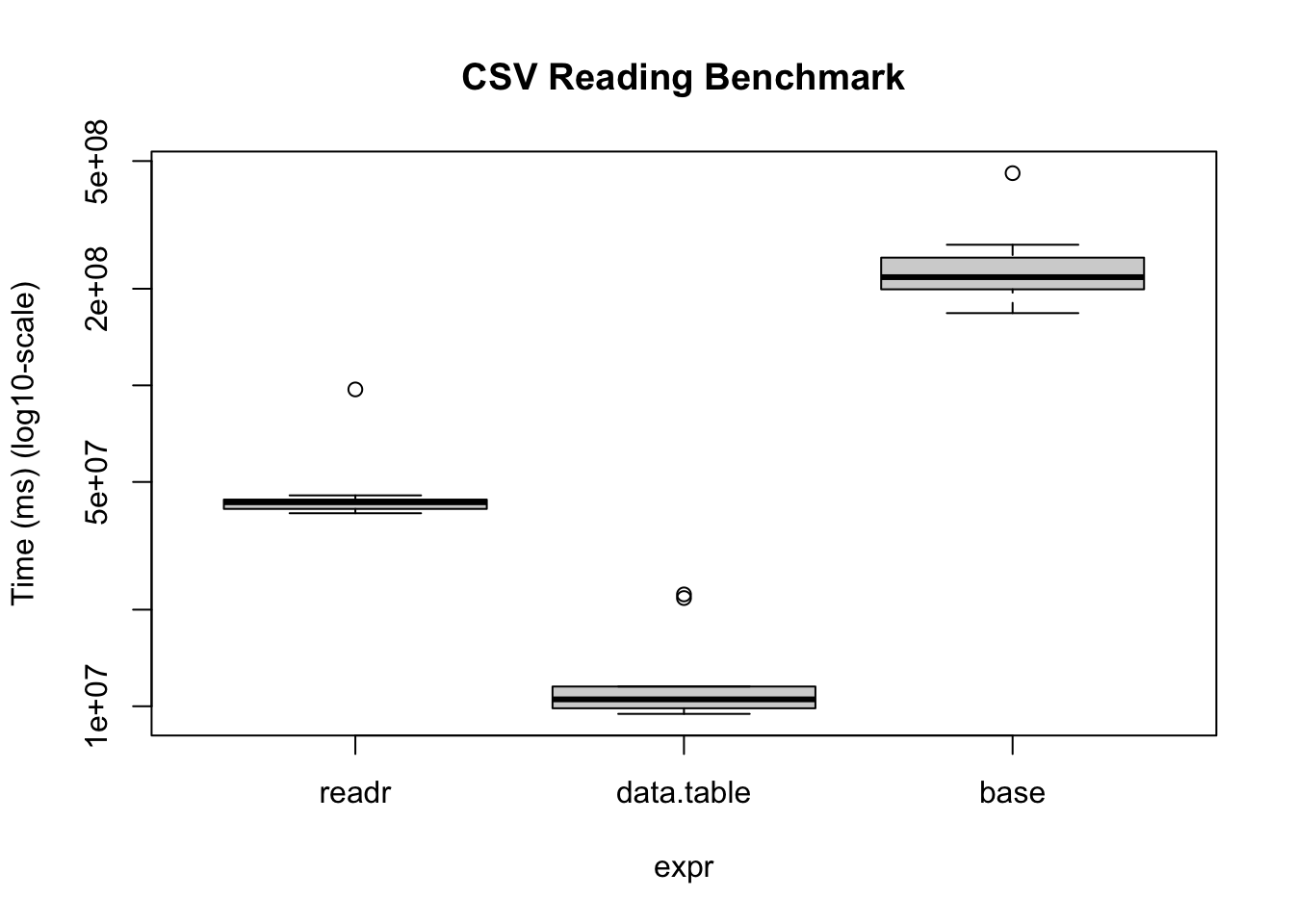
Under the hood, the readr package uses the vroom package. Nonetheless, there are some operations (dealing with character data mostly), where the vroom package shines. Regardless, data.table is a perfect alternative as it goes beyond just reading/writing data.
Example data manipulation
data.table is also the fastest for data manipulation. Here are a couple of examples aggregating data with data table vs dplyr
input <- if (file.exists("flights14.csv")) {
"flights14.csv"
} else {
"https://raw.githubusercontent.com/Rdatatable/data.table/master/vignettes/flights14.csv"
}
# Reading the data
flights_dt <- fread(input)
flights_tb <- read_csv(input, show_col_types = FALSE)library(dplyr)
Attaching package: 'dplyr'The following objects are masked from 'package:data.table':
between, first, lastThe following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, union# To avoid some messaging from the function
options(dplyr.summarise.inform = FALSE)
microbenchmark(
data.table = flights_dt[, .N, by = .(origin, dest)],
dplyr = flights_tb |>
group_by(origin, dest) |>
summarise(n = n())
)Unit: milliseconds
expr min lq mean median uq max neval
data.table 4.451124 4.743495 5.689651 4.931603 5.372681 34.82372 100
dplyr 6.605305 7.039392 7.875169 7.405256 7.844940 22.65771 100